Assembling nanoscale building blocks into an ordered, crystalline structure has attracted significant research interest and has frequently been exploited for developing promising, functional materials. The optimization of inorganic nanoparticle type and size, lattice parameters, and crystallographic symmetry allows for the thermodynamically or kinetically preferred self-assembly of crystalline structures including single, binary, and ternary structures. The emergent properties and diverse chemical functionalities that arise from highly ordered arrangements make them potential platforms for wide applications such as catalysis, magnetics, optics, and functional biomaterials. These developments have inspired similar rules in studies that employ functional protein modules for building tunable periodic arrays with nanoscale precision. However, the chemistry and structural complexity of protein surfaces render it difficult to control their self-assembly into crystals, especially for nonspherical protein modules. Understanding the principle of protein interfacial interactions and thereby organizing anisotropic proteins into protein crystals with tunable structural properties remains a major challenge.
Recently, a research team led by Prof. Qiangbin Wang from Suzhou Institute of Nano-Tech and Nano-Bionics (SINANO), Chinese Academy of Sciences (CAS), demonstrated the controllability of thermodynamics and kinetics in building various periodic superstructures with L-rhamnulose-1-phosphate aldolase (RhuA) modules, by engineering histidine (His)-mediated specific interactions. As shown in Figure 1, RhuA, a C4-symmetric homotetramer, has a square-shaped structure. Based on its unique structural characteristics, a RhuA variant was constructed by replacing Asp47 with His and introducing two His groups at the C terminal (C(2H)), which was denoted by H47RhuAC(2H). The imidazole groups presented two potential interactions: weak π-π stacking interactions and strong metal-histidine chelation. Thus, this strategy could regulate the thermodynamics and kinetics during the assembly processes and construct highly ordered protein crystals with diverse structural properties. For example, H47RhuA-C(2H) molecules were assembled into three-dimensional (3D) nanoribbons and various tetragonal crystals under different conditions. Moreover, this study demonstrated the construction of double-helical superstructures by engineering another His arrangement on the RhuA tetramer surface. Fluorescence microscopy, small-angle X-ray scattering (SAXS), atomic force microscopy (AFM), and transmission electron microscopy (TEM) were employed to characterize the crystalline arrays in detail. These results are expected to pave the way for the better understanding and controlling of periodic protein superstructures with predictable patterns and custom properties.
This work entitled "Noncovalent Self-Assembly of Protein Crystals with Tunable Structures" was recently published in Nano Letters.
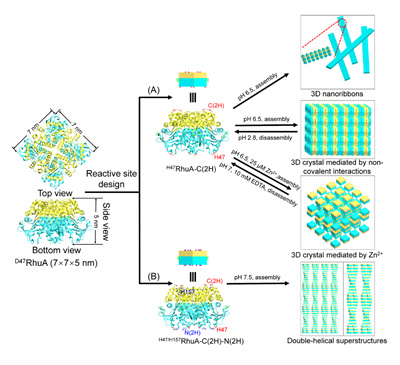
Figure 1. Schematic representation of the formation of 3D nanoribbons, 3D protein crystals, and double-helical protein superstructures. (A) Vertical modification on the top and bottom surfaces of RhuA to obtain a H47RhuA-C(2H) variant. Mediated by noncovalent interactions, H47RhuA-C(2H) variants were self-assembled into 3D nanoribbons and two types of 3D crystals. (B) Design of the H47/H157RhuA-C(2H)-N(2H) variant and the self-assembled, double-helical superstructures via noncovalent interactions.
Contact information: Prof. Qiangbin, Wang, Suzhou Institute of Nano-Tech and Nano-Bionics, Chinese Academy of Sciences
Email: qbwang2008@sinano.ac.cn
Reference: https://dx.doi.org/10.1021/acs.nanolett.0c04587

